Preclinical strategies used to identify potential drug candidates include target-based screening, phenotypic screening, modification of natural substances and biologic-based approaches. In the earlier days of drug de- velopment, phenotypic screening was largely employed and identified molecules that modify a disease phe- noytpe by acting on a previously unidentified target or simultaneously on more than one target. In the 1980s, advances in molecular biology and genomics led to a shift in developing compounds against defined targets that were implicated in disease. The success rate of clinical stage candidates, however has not improved and phenotypic screens are coming back into light.
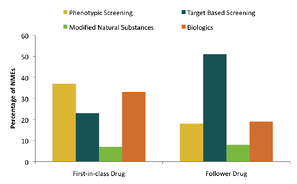 In a recent publication, analysis of different discovery strategies for 259 approved new molecular entities (NMEs) and new biologics between 1999 and 2008 showed that the contribution of phenotypic screening to the drug discovery of first-in-class small molecule drugs exceeded that of target-based approaches in an era when the major focus was on target-based approaches (Swiney, D and Anthony, J. Nature Rev. Drug Disc. (2011) 10:507-519).
In a recent publication, analysis of different discovery strategies for 259 approved new molecular entities (NMEs) and new biologics between 1999 and 2008 showed that the contribution of phenotypic screening to the drug discovery of first-in-class small molecule drugs exceeded that of target-based approaches in an era when the major focus was on target-based approaches (Swiney, D and Anthony, J. Nature Rev. Drug Disc. (2011) 10:507-519).
Phenotypic Screening gets another look?
While phenotypic screening is getting a second look and making its way back into some biopharma's discovery toolbox, many phenotypic screens are established on the basis of cellular systems or systems that are set to ‘mimic’ the in vivo environment. These systems range from a simple single cell types to more complex cell or tissue systems. What can occur with in vitro phenotypic screens that aim to mimic the in vivo environment is:
-
The screen is most often selectively look at single pathways and therefore miss inter-pathway interactions
-
The screens, while aiming to mimic the in vivo environment, don’t predict the in in vivo effect and unexpected biology, interactions or potential toxicity effects that are often observed in vivo.
- Can miss pharmaceutical candidates whose pharmocology isn't evident until it is in a complex biological system.
This has led us to develop a more comprehensive, in vivo phenotypic screening platform, Senerga®. The Senerga® Phenotypic Screening platform consists of a series of matrices designed to cover maximum biological pathways to identify pharmaceutically relevant candidates that also show no predictive toxicity effects early on in the discovery process. This enables researchers to move beyond well-defined targets from the literature or their existing programs and discover new disease biology and potential targets.
The benefit to using this powerful screening program:
- Enables researchers to see effects on disease phenotypes in a complex biological setting with multiple pathways involved.
- Obtain predictive toxicology data at the same time.
- Expands therapeutic potential of libraries as it covers maximum biological pathways with biomarkers to support potential mechanisms
- Identify hits relatively quickly that can be put through further target validation or efficacy proof of concept studies.
With the need to move quickly and fill drug discovery pipelines with new candidates, this phenotypic screening platform is designed to efficiently screen compound libraries rapidly (within 3 - 9 months dependent on the size of the library). The resulting data is mined for hits that are identified from modified disease phenotypes and biological pathways. If you'd like to speak with a scientist about utilizing the Senerga® Phenotypic screening platform, please fill out the following details and a scientist will be in contact with you.







